The era of synthetic biology might finally be upon us. Today, we give you a primer on the latest developments and discuss why we might be near an inflection point for synthetic biology. Note that for this write up, we strive for ease of understanding. As such, we will use analogies and may oversimplify things. The goal is not for you to become a bio-engineer, but rather for you to understand the changing value chain of synthetic biology. Oftentimes, the most critical thing an investor needs to meaningfully evaluate investing opportunities is to understand where different companies and technologies fall in the value chain.
For all the advancement in material science and medicine that mankind has achieved, most of it is accomplished through luck together with trial and error. This is especially true in biology and is largely because we are unable to perform engineering on biology.
The prerequisites for engineering, at its most fundamental level, are the ability to (1) read and (2) write. Only after one can do so in an efficient manner, will repeated iteration of experimentation lead to engineering.
Reading Biology
The first prerequisite of engineering biology is the ability to read DNA.
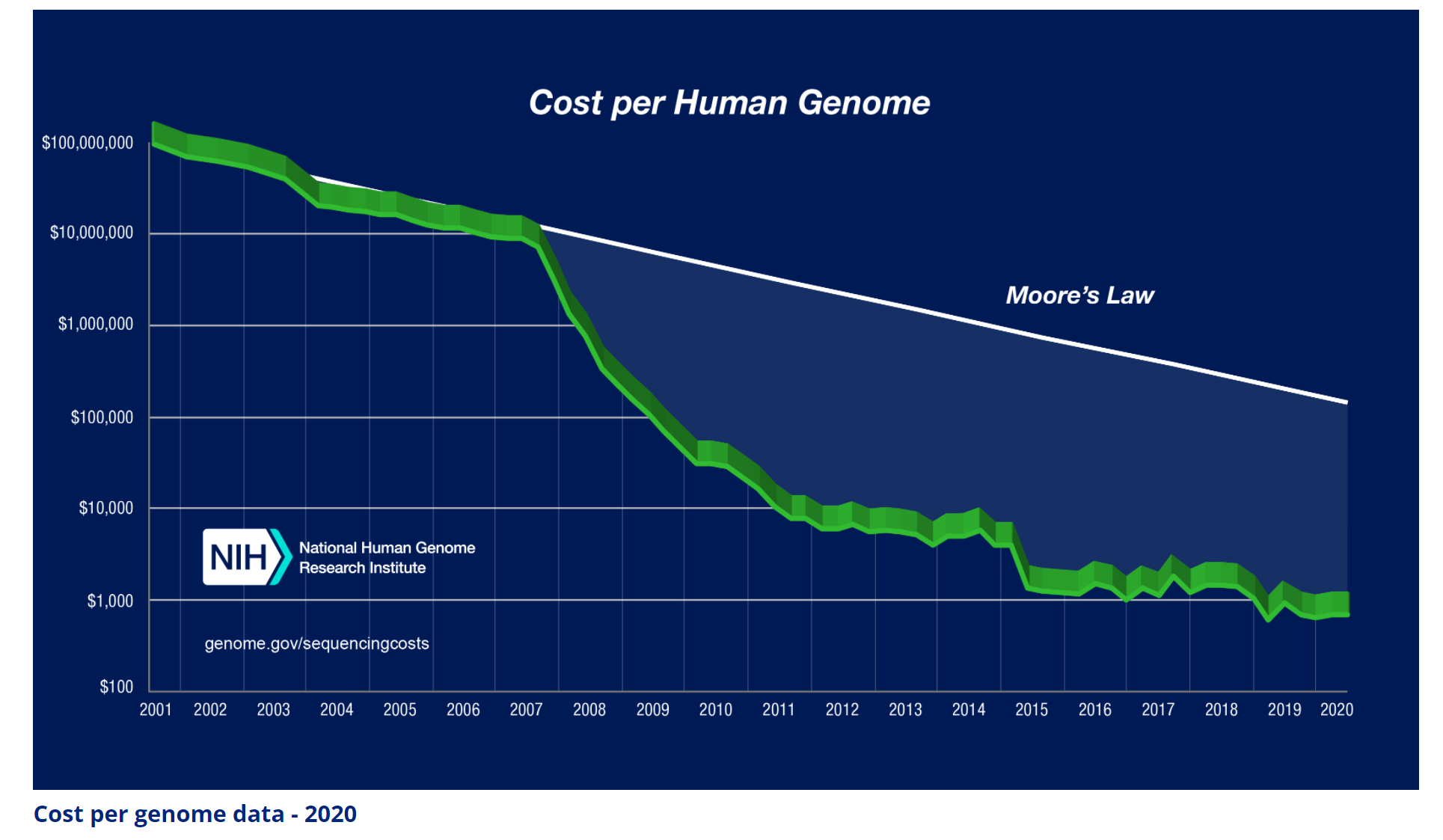
It took 13 years of research and more than $3 billion dollars for the sequencing of the first human genome. But since then, thanks to technology advancements (both in software and hardware), the cost has rapidly declined. So much so that the decline is faster than the decline in transistor cost (Moore’s law). See Figure 1. Nowadays, it costs about $600 and takes about a week to sequence a human’s genome.
Ok, before we go further, we have to explain some jargon:
- The genome consists of all of the DNA contained in a cell’s nucleus.
- DNA is made of four building blocks (A, T, C, G). These are called “basesâ€.
- The information encoded in the DNA is determined by the ordering of these bases.
DNA sequencing is the process where we read the ordering of these bases. With lower costs, the use case of DNA sequencing expanded. Cheaper sequencing allows for:
- Accessible prenatal screening to detect potential genetic disorders.
- Better understanding of why different people react to drugs during drug trials.
- Faster identification of new pathogens. The most recent example of this is the sequencing of the novel coronavirus. The original sequence of the virus was published online on January 11, 2020. Two days later, Moderna announced that it had produced a vaccine.
- Genomic surveillance, the tracking of how the novel coronavirus is constantly changing.
As useful as it has been, however, gene sequencing alone has not translated to the new paradigms and discoveries that many have speculated when it was first announced. For an industry to flourish, the rapidly declining cost must be accompanied by expanding use of the technology. In the case of semiconductors, the decline in transistor cost led to the development of faster and cheaper computers (from large mainframes, to desktop PCs, laptops, and smartphones). Therefore, the market expanded rapidly, leading to the founding of multi-trillion dollar businesses.
In the case of gene sequencing, however, the total addressable market is not expanding as rapidly. It is estimated that the gene sequencing market is worth about $7.5 billion a year and is growing at less than 15% per year. Yes – it’s a healthy growth rate, but it’s nothing spectacular (yet).
Currently, Illumina has the largest market share in the space. It makes sequencing instruments the size of a microwave (or a refrigerator – depending on the product line), with prices ranging from $20,000 to $1 million. A new entrant in the market, Oxford nanopore, has entered the space with a novel technology that enables sequencing at a much faster rate (in hours vs. days with Illumina) and with higher portability (its device is the size of a large smartphone). An increase in speed and portability, along with a decrease in cost, may further expand the sequencing use cases. However, a cheaper cost to read without the ability to write means you cannot create new use cases.
Writing Biology
Before we jump into this section, we need to do two things: (1) understand a bit more about the building blocks of biology and (2) understand why we want to write biology.
The building block of life
Building on our list of jargon:
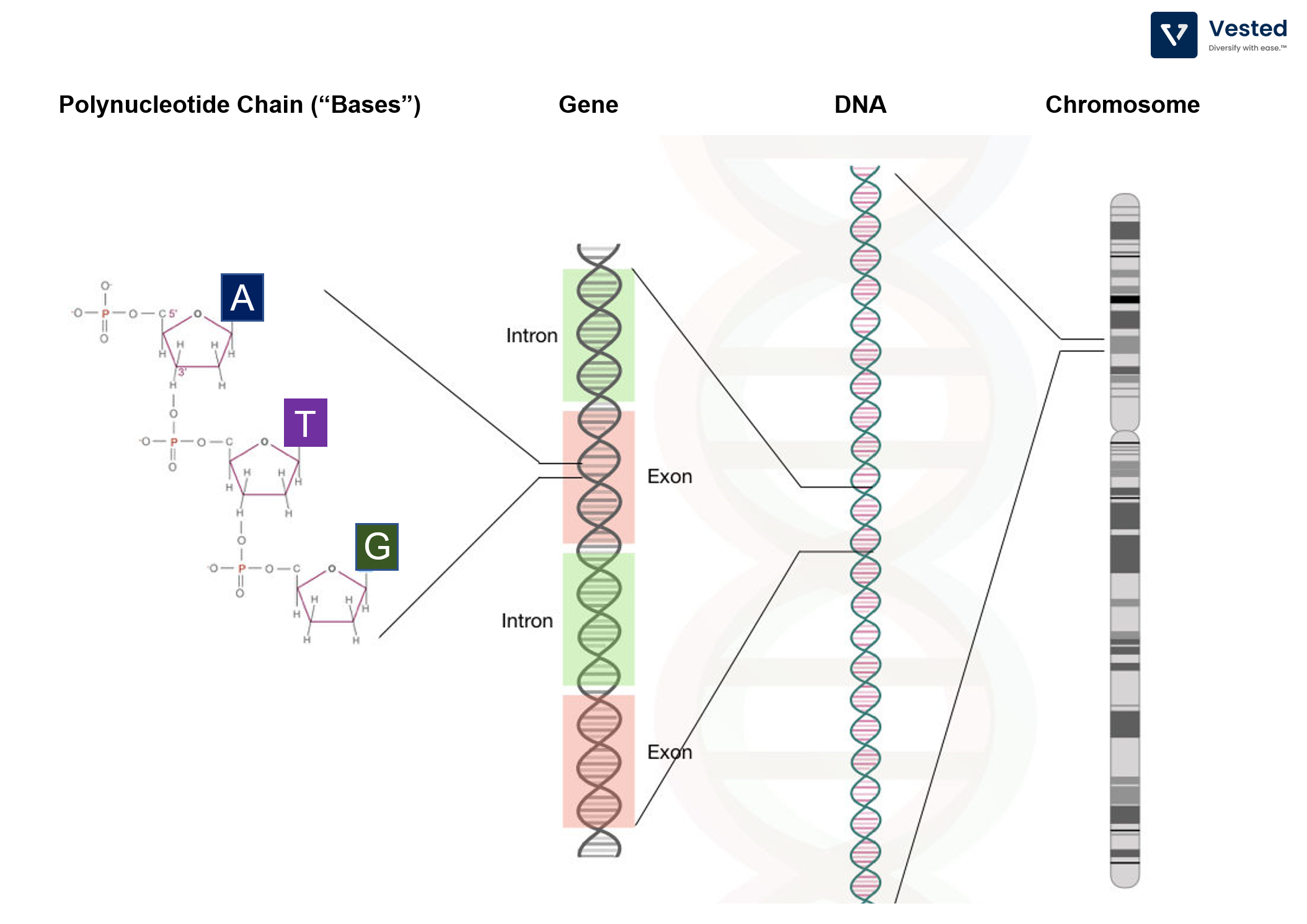
- The bases (A, T, C, G polynucleotide chains) string together to form a strand of DNA. There are typically millions to billions of bases comprising a single strand of DNA. E. Coli bacteria has about 5 million bases and the fruit fly about 123 million bases, while humans have 3 billion bases (of which 99% are the same from one person to another).
- Each segment of DNA has different functions. These functional segments are called genes.
- Most of genetic engineering is directed to modify these functional sections to do what we want – mostly produce specific proteins.
- Proteins do useful things. They can regulate blood sugar (insulin) in the human body or be generated by bacteria to kill pests, among other uses.
Why we want to write biology
Insulin is a type of protein that helps regulate blood sugar levels in the body. In humans and animals, it is produced by the pancreas. In people with diabetes, their ability to produce insulin is hampered. Before Genentech came about (founded in 1976), insulin was produced by harvesting animal pancreas. The process is very inefficient and expensive. At the time, it took 8,000 pounds of pancreas glands from 23,500 animals to make one pound of insulin. By the late 70s, Eli Lilly, the main producer of insulin at the time, needed more than 56 million animals annually to meet demand.
Then came Genentech, the first biotech company. The company was able to modify a specific gene in E. Coli bacteria – enabling the bacteria to produce insulin, which was then harvested for human use. This significantly lowered the cost of insulin and expanded access to the medication to more people.
As you can see, writing DNA gives us the ability to use biology as our manufacturing tools. This can have profound implications in creating new medicine or new materials. However, while the process of writing DNAhas been around since the 80s, it hasn’t become significantly cheaper (Figure 3).
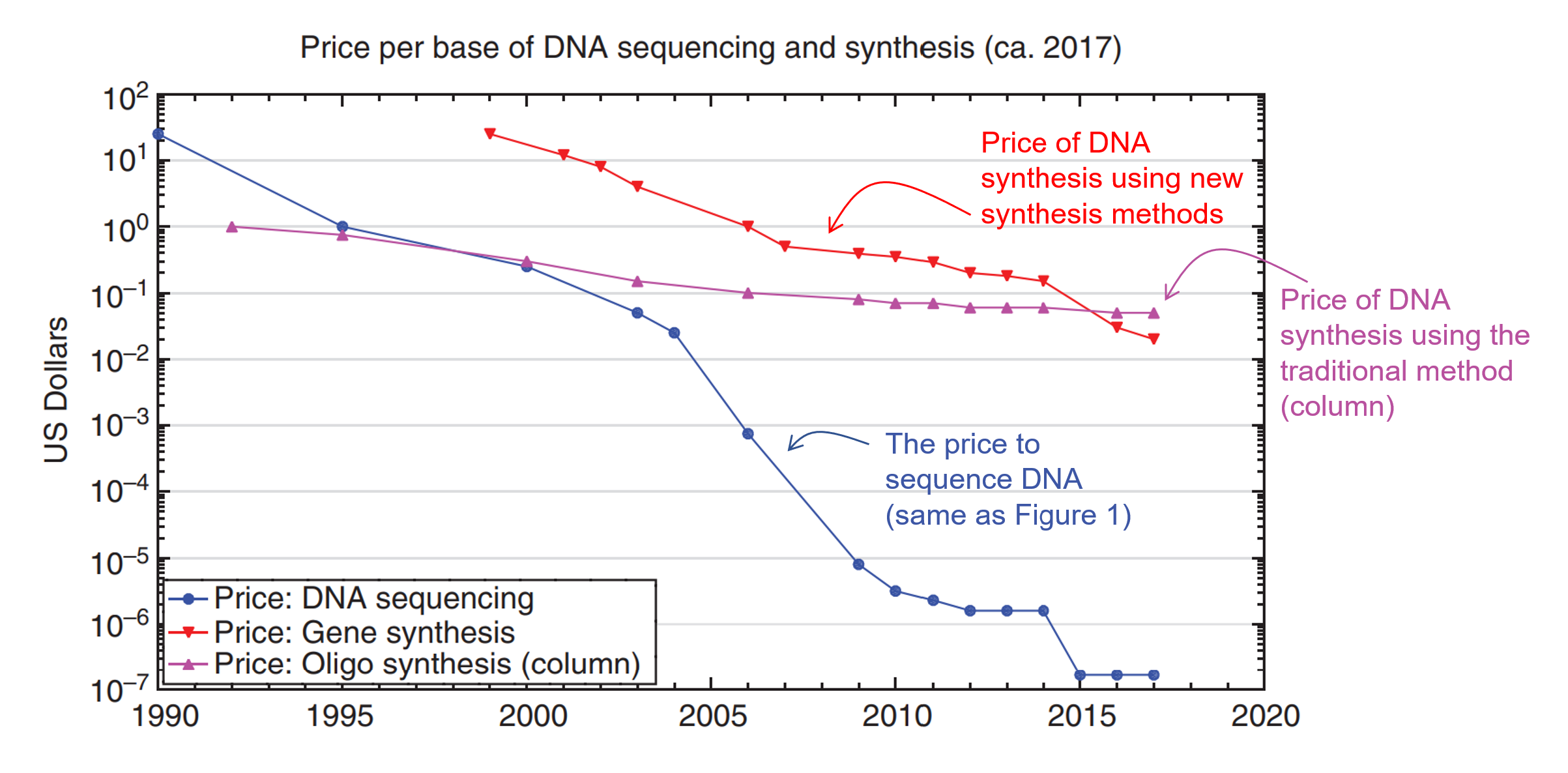
As you can see from Figure 3, the cost decline for DNA synthesis has not kept pace with the decrease in cost for DNA sequencing (reading). From the 90s to the late 2020s, DNA sequencing costs went down by a factor of 10 million times (blue line – this curve is called the Carlson Curve). But the traditional cost of writing DNA (purple line) only went down by 100x. It is only in recent years that the cost of making DNA using alternative methods (red line) crossed the threshold where it is now cheaper than the traditional method.
This is why so many people are excited that a new age of synthetic biology might be upon us. As you can see, the red curve is trending down faster than the purple line.
A new way to write biology
The traditional way
The traditional method of writing DNA is using the Phosphoramidite Oligonucleotide Synthesis method. It’s basically an assembly process that assembles one base at a time, stitching either A, T, C, or G at a time, in the correct order. And traditionally, the assembly process is done with pipettes, in a plastic well, where each well produces only one sequence of genes.
The process is time consuming and expensive (requires a lot of high tech equipment, skilled labor, and reagents). So, how can we improve throughput and lower costs? We can make the well as small as possible.
The new way – writing from scratch
One way to do that is to leverage semiconductor manufacturing methods (which are suitable to work at nanoscale) and create microscopic wells on silicon wafers, such that each processing step acts on 9,600 wells, rather than 1 well – effectively increasing throughput by 9,600x. That is the process pioneered by Twist Bioscience (which went public in 2018). Figure 4 below shows the comparison of the traditional assembly process.
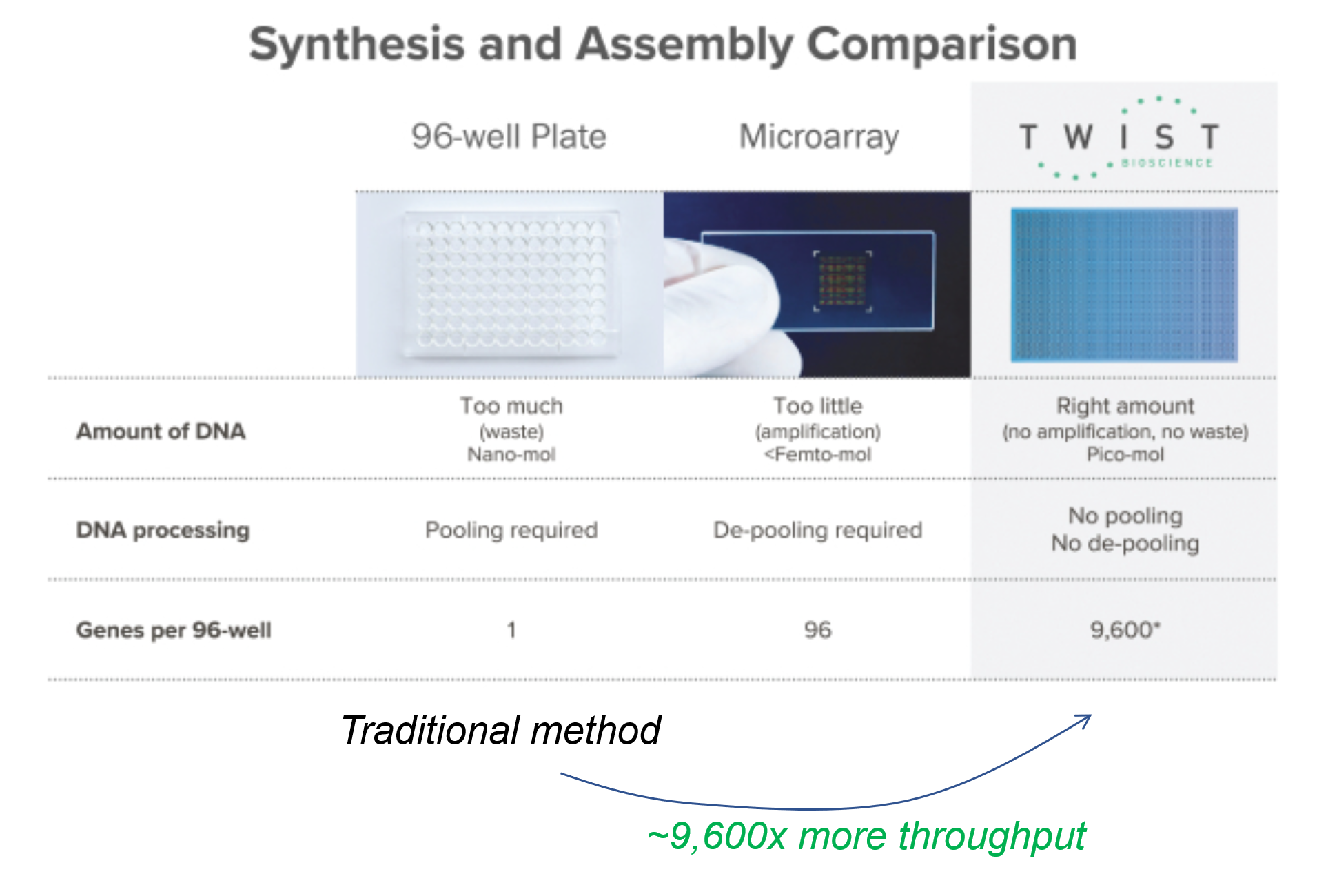
Twist’s business is that of a DNA manufacturing service. Customers can order novel sequences from the company, or they can integrate its API into their own software. In a sense, Twist is building a B2B infrastructure to write custom DNA at scale – similar to what AWS pioneered in cloud computing.
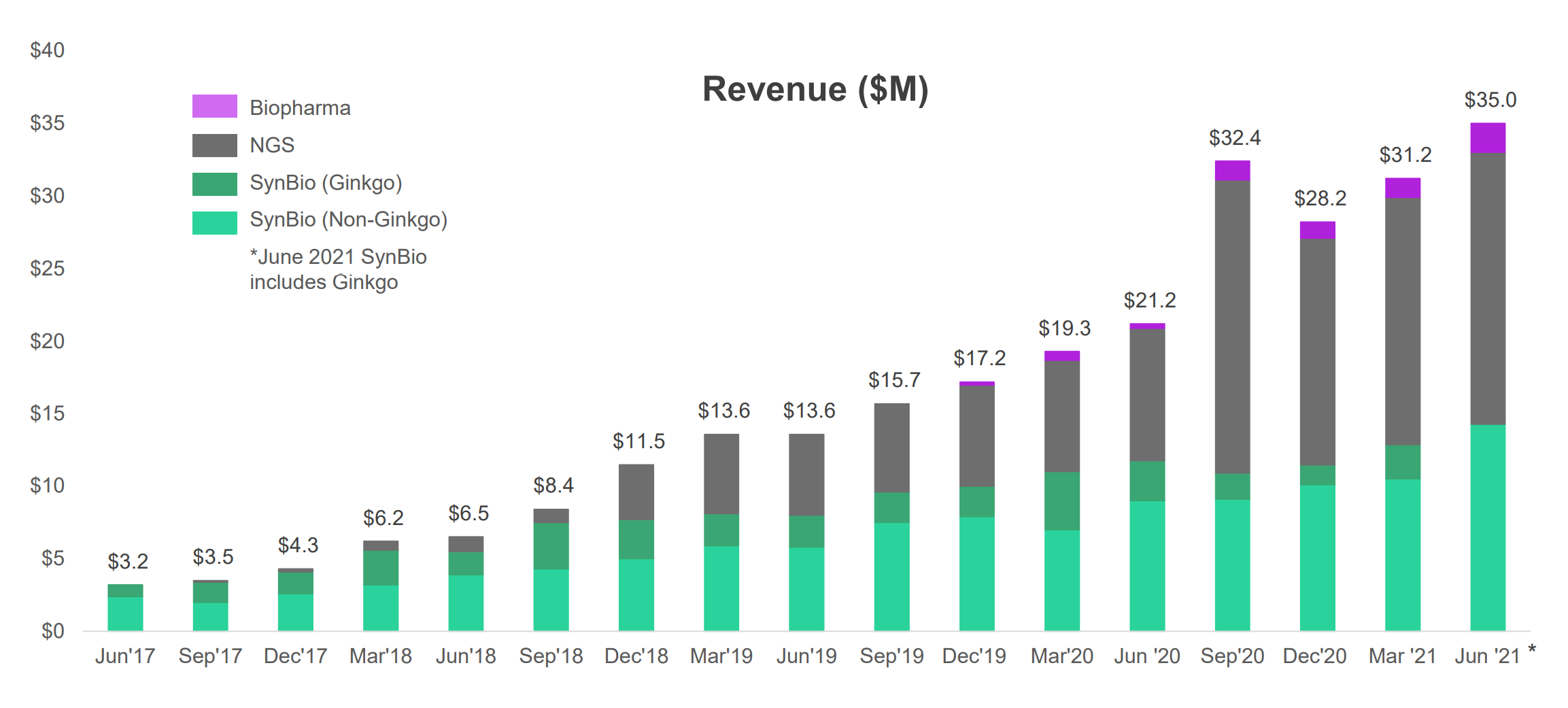
Sounds exciting? Sure does. But hold on! Since its IPO, Twist has been able to grow its revenue rapidly, but despite that, the share price has largely been down this year (about half of its peak from January 2021). Clearly, the valuation far exceeded reality and has since been re-assessed by the market. Even at this share price point, the valuation is still at 32.5x next twelve month price-to-sale ratio (as a comparison, Robinhood is about half of this).
So far, we’ve discussed ways to write DNA sequences from scratch. But another way to write is by copy-paste-edit method. That’s where CRISPR comes in.
Another way – cut and edit
CRISPR-Cas9 is akin to a homing missile that can target a specific genetic sequence and cut said sequence. A broken genetic sequence can then be terminated (useful if you have an undesirable mutation) or can be edited (by inserting a new sequence) to do something useful. This method is inspired by the immune system of bacteria (when fighting viruses).
This way, we don’t have to write the genetic sequences from scratch, which is hard to do. The discovery of CRISPR resulted in a Nobel prize in Chemistry in 2020 and a heated patent litigation that has yet to be resolved. But what is interesting to investors is that the technology has been de-risked enough that there are a slew of companies employing this technology to treat various diseases (see Table 1).
Table 1: Publicly traded CRISPR companies
| Ticker | Company | Description |
| NTLA | Intellia Therapeutics | CRISPR-Cas9 gene editing to treat protein disorders |
| CRSP | CRISPR Therapeutics | CRISPR-Cas9 gene editing to treat blood disorders |
| BEAM | Beam Therapeutics | Developing gene editing to treat blood and ocular diseases, and for oncology purposes |
| EDIT | Editas Medicine | Developing base editing to treat blood, liver, and ocular diseases |
Notice that all these CRISPR companies are focused on treating genetic disorders. This is because they are editing, not writing. Knowing a reference sequence for the human genome, they are mostly focused on repairing the sequence to restore functionality. On the other hand, companies that are trying to write new sequences are largely trying to get functionalities to produce proteins in a specific way.
Intelligent design
Just because one can arrange letters in a specific sequence, does not mean one can craft a prose. Just because we can arrange genetic sequences in the order that we want and in a low cost manner, does not mean we can get the functionality that we desire.
To help cell programmers achieve desired results, synthetic bio design companies, such as Ginkgo Bioworks (who just SPACed) are building platforms (both software and hardware) to discover new genetic sequences that work. Ginkgo buys synthetic DNA from Twist (see Figure 5, Ginkgo is one of Twist’s biggest customers) and runs experimentation on its own platform.
Since we do not have an accurate predictive model for biology, the design stage for the most part means throwing stuff at the wall (intelligently) and seeing what sticks. This means, increasing experimentation throughput is key – and that is what Ginkgo’s platform does. Their platform has two core assets: Foundry and Codebase.
- The Foundry is a combination of software and hardware (robots) that stitch together cell engineering workflows: designing DNA, writing DNA, inserting DNA into cells, testing performance, and carrying out analytics. When Gingko first built the Foundry, it was less efficient than a skilled scientist in the lab. But since 2015, Ginkgo has been able to reduce cost per unit operation by ~50% every year.
- The Codebase is a repository of intellectual property (genetics parts, sequences, performance data) that the company leverages for future designs. As Ginkgo carries out more experiments, this repository grows.
Figure 6 below illustrates how Ginkgo’s platform helps its customers discover new biology. Notice the high number of synthetic base pairs evaluated!
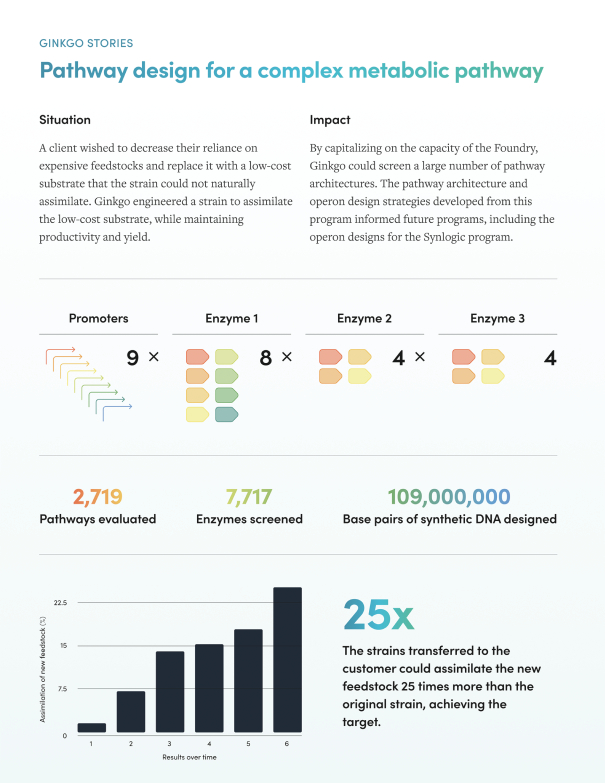
As a company that has recently SPACed and is doing something unique in a hot market (synthetic biology), Ginkgo’s valuation is expensive. At 127x next-twelve-month price-to-sale ratio, it is valued 4x higher than even Twist.
Closing Thoughts
Throughout this article, we make multiple analogies to the semiconductor industry. But there are two critical differences:
First, lower costs is a prerequisite but not a guarantee of greater demand. There is still a big question if the market can expand fast enough. Intel’s and TSMC’s business model of incessantly lowering the cost of transistors works because the growth of the market far exceeded the decline in cost of transistors.
But in the case of synthetic DNA, if the demand does not expand at a rate that is faster than the rate of decline in cost of DNA sequencing, revenue of the synthetic DNA producers will shrink. This is why Twist needs companies such as Ginkgo to help it generate more demand.
Second, in the semiconductor industry, building that first chipset at the leading technology node is only the first step to actually building a competitive moat around the process. It still takes years of R&D and $10-15 billion in capital expenditures to build a factory that is capable of producing said technology nodes at sufficient scale, yield, and productivity.
In other words, in the semiconductor industry, being at the leading edge means that you can deliver the lowest cost transistor and have the manufacturing capacity to deliver said transistor (both cost and process moat).
In contrast, in synthetic DNA, the hard part is to design and produce that first strand of DNA. After that, you can photocopy the novel DNA in a very cheap way, using PCR (a technique that’s been around since 1983). In other words, the need to write a specific synthetic DNA stops the moment you have validated that the synthetic DNA works (because after that, you can photocopy the DNA using PCR). This means two things: (1) to continue to sustain the demand of writing synthetic DNA, the increase of demand has to be even greater than similar dynamics in semiconductors, and (2) the technology moat may not exist. The competitive moat might be from lower cost and throughput, but not from a differentiated process.